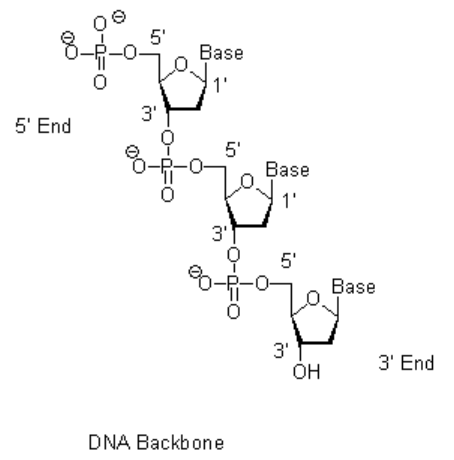
The conformation of the sugar-phosphate backbone accompanying intercalation has been accurately determined in this analysis, and contains the mixed sugar-puckering pattern: C3' endo (3'-5') C2' endo. The other ethidium molecule stacks on either side of the intercalated base-paired dinucleoside monophosphate, being related by a unit cell translation along the a axis. Base-pairs in this duplex are separated by 6.7 A this reflects intercalative binding by one of the ethidium molecules. The two UPa molecules are hydrogen-bonded together by Watson-Crick base more » pairing. The asymmetric unit consists of two ethidium molecules, two UpA molecules and 19 solvent molecules, a total of 145 non-hydrogen atoms. This light atom structure has been solved to atomic resolution and refined by full matrix least squares to a residual of 0.12, using 3034 observed reflections. The complex crystallizes in the monoclinic space group P2/sub 1/ with unit cell dimensions, a = 13.704 A, b = 31.674 A, c = 15.131 A. « lessĮthidium forms a crystalline complex with the dinucleoside monophosphate, uridylyl (3'-5') adenosine (UpA). 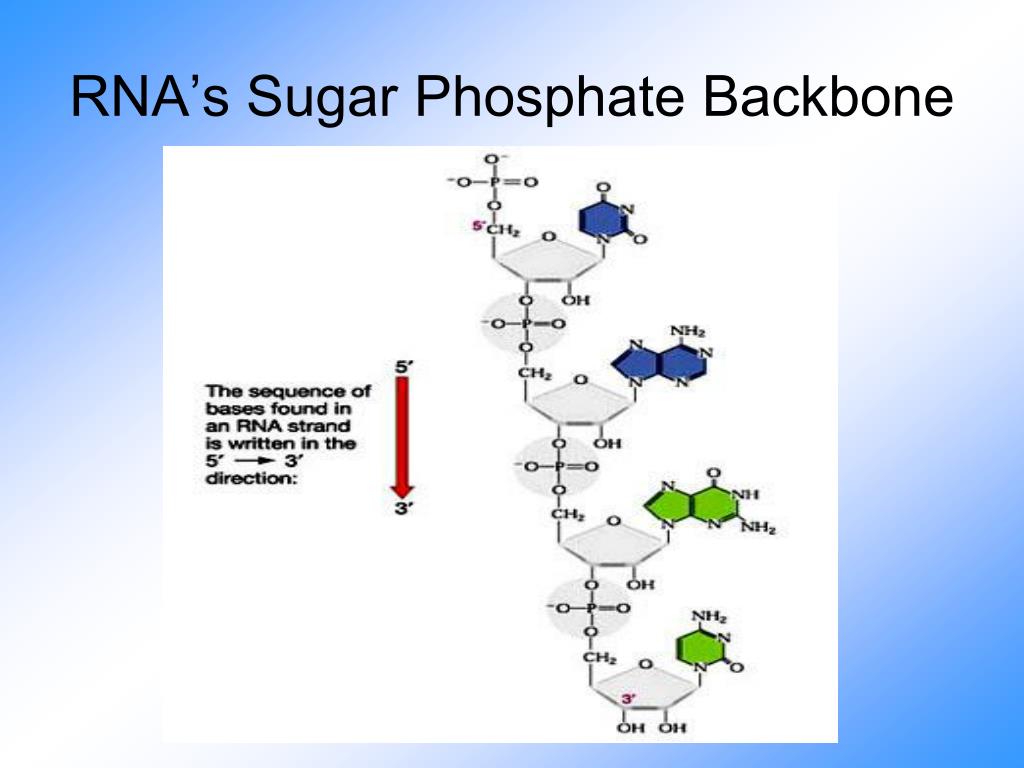
Our structure offers the most detailed view to date of DNA solvation in the solid-state undistorted by metal ions or polyamines. Exocyclic amino groups of guanine (N2) and cytosine (N4) unexpectedly stabilize waters H-bonded to O2 keto oxygens from adjacent cytosines and O6 keto oxygens from adjacent guanines, respectively. For example, a water molecule located at H-bonding distance from O6 keto oxygen atoms of two adjacent guanines directs both its deuterium atoms away from the keto groups. Several water molecules display entirely unexpected orientations. In particular, H-bond acceptor and donor patterns for water participating in prominent hydration motifs inside the minor groove, on the convex more » surface or bridging nucleobase and phosphate oxygen atoms are finally revealed. The neutron density visualized at 1.5 Å resolution for the first time allows us to pinpoint the orientations of most of the water molecules directly contacting the DNA and of many second-shell waters. We used cryo neutron crystallography to gain insight into the H-bonding patterns of water molecules around the left-handed Z-DNA duplex 2. « lessĮven in high-quality X-ray crystal structures of oligonucleotides determined at a resolution of 1 Å or higher, the orientations of first-shell water molecules remain unclear. geometry, conformation, protonation state and hydration more » structure. We demonstrate that joint cryo-neutron/X-ray crystallography offers an all-in-one approach to determine the complete structural properties of RNA, i.e. We discuss 2'-OH and water molecule orientations in the SRL neutron structure and compare with results from a solution phase 10 μs MD simulation. Neutrons reveal the strong preference of the 2'-OH to donate in H-bonds and that 2'-OH orientation affects both backbone geometry and ribose pucker. An atypical C2'-endo sugar pucker is strictly correlated with a 2'-OD sugar-orientation. GAGA-tetraloop riboses display a 2'-OD base-orientation. Most riboses in the SRL stem region show a 2'-OD backbone-orientation. 
2'-OD orientations were established for all 27 residues and revealed O-D bonds pointing toward backbone (O3', 13 observations), nucleobase (11) or sugar (3). Here, we report the first neutron crystal structure of an RNA, the Escherichia coli rRNA Sarcin-Ricin Loop (SRL). Thus, standard crystallography cannot determine 2'-OH orientation (H2'-C2'-O2'-HO2' torsion angle) and its potential roles in sculpting the RNA backbone and the expansive fold space. Macromolecular X-ray diffraction experiments typically do not reveal the positions of hydrogen atoms. The ribose 2'-hydroxyl is the key chemical difference between RNA and DNA and primary source of their divergent structural and functional characteristics.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
